Passbands Tutorial
This tutorial shows how to create and use passband objects with response functions that can be (1) tabulated from files, or (2) have top-hat shapes.
Note
You can also find this tutorial as a Python scrip here or as a jupyter notebook here.
Let’s start by importing some necessary modules:
import pyratbay.io as io
import pyratbay.spectrum as ps
import pyratbay.constants as pc
import matplotlib.pyplot as plt
import numpy as np
1. Passband objects
1.1 Passband from tabulated file
Passband objects can be created by reading from a plain text file that tabulates the wavelength (in microns, first column) and its response function (second column), e.g.:
filter_file = f'{pc.ROOT}pyratbay/data/filters/spitzer_irac2_sa.dat'
band = ps.PassBand(filter_file)
Display some useful data that is contained in the object:
print(f'Filter name: {band.name}')
print(f'Central wavelength: {band.wl0:.3f} um\n')
print(f'Filter file: {band.filter_file}\n')
print(f'Wavelength array (um): {band.wl}')
print(f'Wavenumber array (cm-1): {band.wn}')
print(f'Response function: {band.response}')
Filter name: spitzer_irac2_sa
Central wavelength: 4.471 um
Filter file: /Users/pato/Dropbox/IWF/projects/2014_pyratbay/pyratbay/pyratbay/data/filters/spitzer_irac2_sa.dat
Wavelength array (um): [5.222 5.2167 5.2115 ... 3.7278 3.7252 3.7225]
Wavenumber array (cm-1): [1914.9824 1916.9133 1918.8407 ... 2682.5186 2684.4485 2686.3739]
Response function: [0.0004 0.0004 0.0004 ... 0.0007 0.0007 0.0008]
Evaluate passband over a specific wavelength array (um):
wl = np.arange(3.5, 5.5, 0.001)
out_wl, out_response = band(wl)
# Show profile:
plt.figure(1)
plt.clf()
plt.plot(out_wl, out_response, color='darkorange', lw=2.0)
plt.axhline(0.0, color='gray', lw=1.0, dashes=(10,2))
plt.xlabel('Wavelength (um)')
plt.ylabel('Response function')
# Note that returned values are the same as the internal variables:
# out_wl = band.wl
# out_response = band.response
# The variable band.idx points to the indices in input wl
# that overlap with the band's wavelength:
# (Note wl differs from band.wl)
plt.plot(wl[band.idx], band.response, color='blue', lw=1.0, dashes=(5,5))
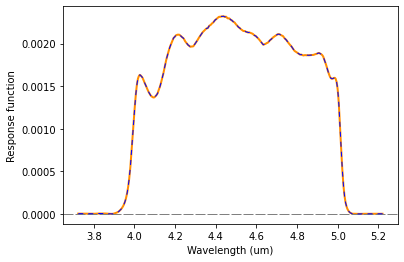
The band response function is scaled such that the integral over wavenumber equals one:
band_integral = np.trapz(band.response, band.wn)
print(f'Integral of response function: {band_integral:.3f}')
Integral of response function: 1.000
It is possible to evaluate the passband over a wavenumber array:
wn = 1e4 / wl
out_wn, out_response = band(wn=wn)
plt.figure(1)
plt.clf()
plt.plot(band.wn, band.response, color='black')
plt.xlabel(r'Wavenumber (cm$^{-1}$)')
plt.ylabel('Response function')
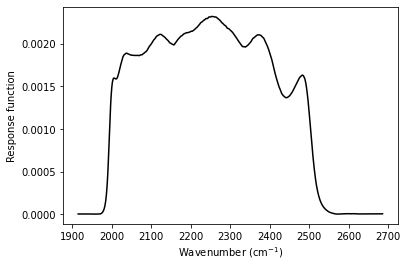
1.2 Top-hat passband
Top-hat passband objects can be created by setting their central wavelength and half-width (in micron units):
wl0 = 4.5
half_width = 0.5
band = ps.Tophat(wl0, half_width)
# Evaluate passband over a specific wavelength array (um):
wl = np.arange(3.5, 5.5, 0.001)
out_wl, out_response = band(wl)
# Show profile:
plt.figure(1)
plt.clf()
plt.plot(out_wl, out_response, color='black', lw=2.0)
plt.xlim(3.7, 5.3)
plt.axhline(0.0, color='gray', lw=1.0, dashes=(8,2))
plt.xlabel('Wavelength (um)')
plt.ylabel('Response function')
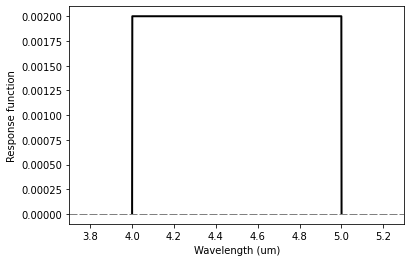
Same as with ps.Passband(), a tophat response function is scaled
such that the integral over wavenumber equals one:
band_integral = np.trapz(band.response, band.wn)
print(f'Integral of response function: {band_integral:.3f}')
Integral of response function: 1.000
2. Create a set of passbands
If you have a set of observations for a given target, you can generate a set of filters for these data by loading the info from a file like the one below (not how you can combine top-hat and from filter files):
# observations.dat file
# Passband info could be (1) a path to a file or (2) a tophat filter
# defined by a central wavelength, half-width, and optionally a name
# Comment lines (like this one) and blank lines are ignored,
# central-wavelength and half-width units are always microns
# @DEPTH_UNITS sets the depth and uncert units (none, percent, ppt, ppm)
# and also indicates that there's data and uncerts to read
# as two columns before the passband info
@DEPTH_UNITS
ppm
@DATA
# depth uncert wavelength half_width passband_name
# depth uncert passband_file
200 50 0.900 0.075
214 82 1.148 0.046 HST_WFC3
325 83 1.240 0.046 HST_WFC3
415 82 1.332 0.046 HST_WFC3
621 97 1.424 0.046 HST_WFC3
765 101 1.516 0.046 HST_WFC3
732 107 1.608 0.046 HST_WFC3
1148 84 {ROOT}/pyratbay/data/filters/spitzer_irac1_sa.dat
1275 92 {ROOT}/pyratbay/data/filters/spitzer_irac2_sa.dat
Load this data and print a brief summary to screen:
# Loading:
obs_file = 'observations.dat'
bands, depths, uncerts = io.read_observations(obs_file)
# Evaluate pass band over a given spectral array:
wl = np.linspace(0.5, 5.5, 1000)
for band in bands:
band(wl)
# Summary of data:
print('band wl (um) depth (ppm)')
for i,band in enumerate(bands):
depth = depths[i] / pc.ppm
err = uncerts[i] / pc.ppm
print(f'{band.name:16s} {band.wl0:.2f} {depth:9.1f} +/- {err:5.1f}')
band wl (um) depth (ppm)
tophat 0.90 200.0 +/- 50.0
HST_WFC3 1.15 214.0 +/- 82.0
HST_WFC3 1.24 325.0 +/- 83.0
HST_WFC3 1.33 415.0 +/- 82.0
HST_WFC3 1.42 621.0 +/- 97.0
HST_WFC3 1.52 765.0 +/- 101.0
HST_WFC3 1.61 732.0 +/- 107.0
spitzer_irac1_sa 3.52 1148.0 +/- 84.0
spitzer_irac2_sa 4.47 1275.0 +/- 92.0
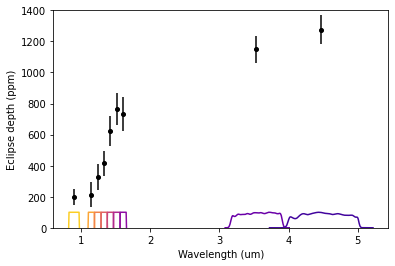
Plot the data and passbands:
nbands = len(bands)
colors = [plt.cm.plasma_r(0.1 + 0.1*i) for i in range(nbands)]
bands_wl = [band.wl0 for band in bands]
plt.figure(10)
plt.clf()
plt.errorbar(bands_wl, depths/pc.ppm, uncerts/pc.ppm, fmt='ok', ms=4.0)
for i,band in enumerate(bands):
response = band.response / np.amax(band.response) * 100
plt.plot(band.wl, response, color=colors[i])
plt.ylim(0, 1400)
plt.xlabel('Wavelength (um)')
plt.ylabel('Eclipse depth (ppm)')